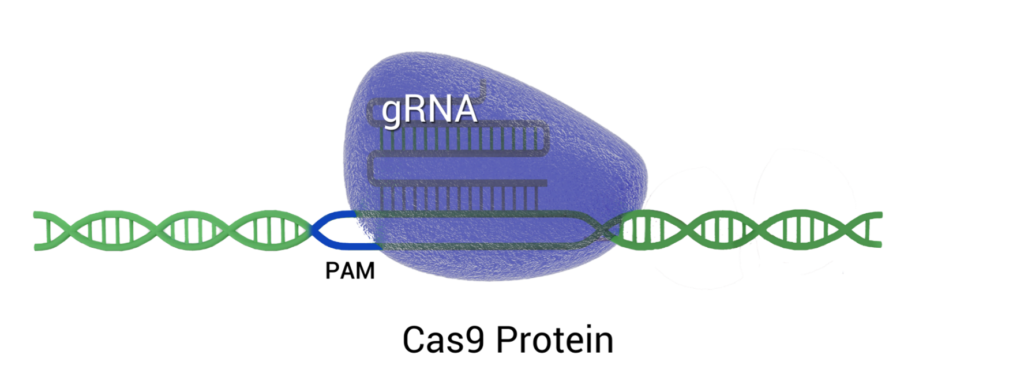
CRISPR/Cas9 uses a single guide RNA(gRNA) and the Cas9 nuclease to bind and cut DNA, enabling targeted genome edits through blunt-ended double-strand breaks. Repairs occur mainly through Non-Homologous End Joining (NHEJ), creating insertions or deletions at the cut site.
Because Cas9 is monomeric and relies on a single gRNA, the system is known to generate unintended off-target edits and genomic rearrangements. In addition, CRISPR/Cas9 lacks a clear freedom-to operate (FTO) landscape for many commercial applications.