Optimized Agricultural Gene Editing
Advanced gene editing enables transgene-free targeted plant traits to support breeding programs.

Enabling Crop Engineering With Seamless Gene Editing
Traditional breeding techniques are time-consuming, while transgenic methods result in the presence of exogenous genes, leading to “GMO” labels and substantial regulatory expenses.
Modern CRISPR based gene editing tools such as Cas-CLOVER enable crop breeders to perform precise genetic modifications without introducing foreign genes, thereby producing non-GMO crops. This scarless gene editing method is essential for addressing the increasing demands for food, feed, and energy.
Cas-CLOVER More Precise Than CRISPR/Cas9
The CRISPR/Cas9 system has been proven to introduce numerous off-target mutations, potentially unintentionally changing germplasm and introducing regulatory risk. Consequently, backcrossing is necessary to eliminate these undesired mutations.
Cas-CLOVER, a high-fidelity dimeric nuclease, facilitates precise gene knockouts with reduced off-target effects. This advanced CRISPR based system is an invaluable asset for trait discovery and introduction initiatives.
Cas-CLOVER is Validated for Plant Gene Editing
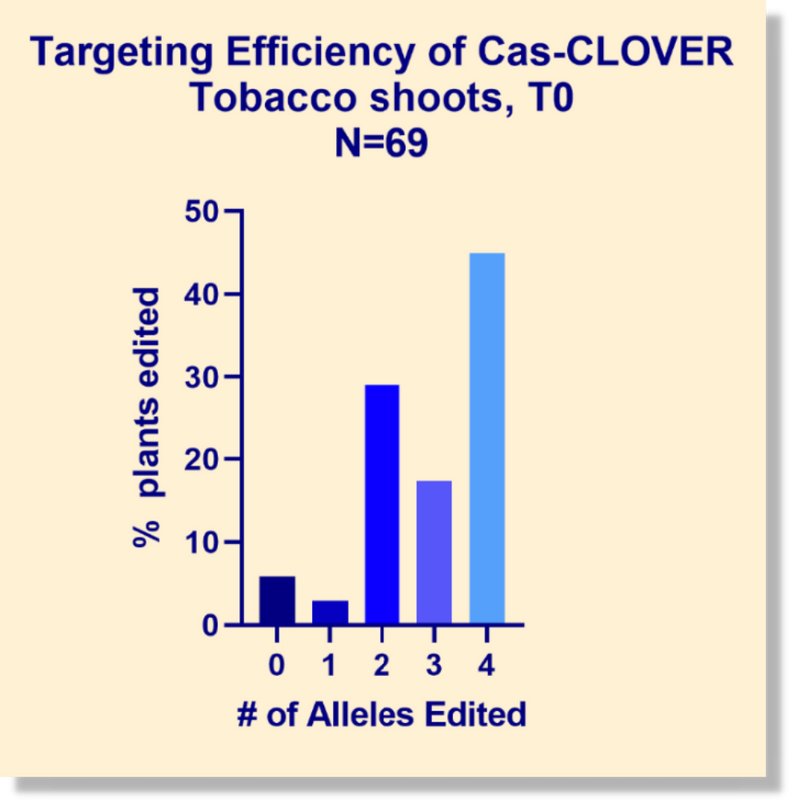
Cas-CLOVER was validated in plants by generating knockout mutations in both monocot and dicot species. In our laboratory, we targeted phytoene desaturase (PDS) and dozens of other genes in the crop research model, tobacco. The PDS knockouts resulted in the anticipated albino phenotype. Screening indicated that nearly 50% of the regenerated shoots exhibited edits in all four alleles, demonstrating Cas-CLOVER’s high efficiency.
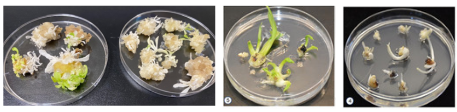
Displayed are albino phenotypes yielded in tobacco and banana via Cas-CLOVER knockouts. From left:
- Tobacco callus and shoots edited with Cas-CLOVER
- Control non-edited banana shoots.
- Banana embryo germinated from transformed embryogenic cell suspension (ECS)
PiggyBac Validated In Plants For Trait Enhancement
As in most plants, targeted homologous recombination (HR) mediated precision genome editing in rice is an infrequent event. Efficiency can be vastly improved by positive-negative selection. However, the positive selection marker needs to be seamlessly removed from the locus following editing.
Small to very large stable gene integration (200KB+) enables whole pathway metabolic engineering and high expression of proteins. In this example, we integrated the Mevalonate Pathway (approximate 30 kb) with piggyBac and saw an almost 9-fold increase in β-Farnesene production with no optimization of growth conditions.
See How Scientists Are Using Our Gene Editing Tools

Principal Scientist In Plant Biotechnology At The International Institute of Tropical Agriculture
“We are very excited to improve the banana varieties preferred by farmers in Africa for disease-resistant bananas using Cas-CLOVER. The gene-edited banana with no foreign gene integration will not be regulated as a GMO in Kenya. The disease-resistant varieties will enhance banana production and increase the incomes of smallholder farmers in East Africa, where the banana is one of the major staple foods.”
Optimizing Your AgBiotech Research?
Contact us to learn more about our innovative gene editing technology.
