In recent years, some of the biggest advances in agriculture and health have stemmed from the development of genetically engineered foods. Research teams have increased their efforts in searching for new ways of inserting exogenous genes into plant DNA for their specific traits. So far genetic engineering has shown plenty of promise and is constantly changing the food we eat. Since the mid-1990’s, a wide variety of foods derived from engineered ingredients have been on the shelf. Well over half of all processed foods in U.S. supermarkets contain at least one type of engineered soybeans, corn, or canola. This includes everything from pizza and chips, to cookies, corn syrup, and even baking powder. One good example of this is the Lycopersicon, also known as the wild tomato. Genetic engineering has enabled rapid enhancement of the fruit from the size of a marble into gigantic beefsteak tomatoes. In fact, over 50 different designer crops have been federally approved, with about a hundred more undergoing further research.
There are myriad reasons to cultivate genetically engineered plants, which are created by inserting a gene or genes from a donor organism carrying desired traits into the host organism in order to express the desired trait. Nowadays, researchers are able to pull a desired gene from just about any species and insert it into another organism. For example, genes can be inserted in order to make a plant produce vitamin C, or splice genes in for protection from bacterial disease as well as introduce a vaccine for distribution especially in economically disadvantaged sections of the globe.
On a genetic level, humans have been altering crops for millennia. By choosing specific traits and breeding only those plants one selects for certain genetic changes, effectively engineering the crop. One key advantage of selective breeding is that no exogenous DNA or transgenes are introduced into the plant that will be consumed. However, selective breeding for traits takes decades and the outcome is uncertain. What if we could make the same edits as selective breeding, without adding transgenes, but faster with advanced technology? Enter seamless gene editing.
It’s known that close to 1 billion people on the planet are actually malnourished, and the number will continue to grow. Seamless gene editing could be a huge solution to problems of food shortages and hunger, because it can increase crop yields, offer more varieties of products, and help plants resist pests and diseases even better than before.1
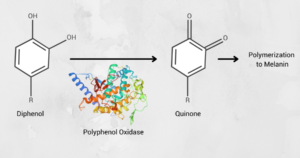
Figure 1: Polyphenol oxidases are necessary for the production of melanin, which give fruit a brown color
Reducing Enzymatic Browning In Potato Tubers
One of the biggest hurdles for potato farmers is something called enzymatic browning, and it’s a serious problem for everyone including producers, distributors, and even the consumers. In the potato (Solanum tuberosum L.), enzymatic browning can have devastating effects during harvest and post-harvest procedures such as shipping, storage, and during distribution to stores. Currently this undesired process is controlled only by chemicals, which is not a healthy or environmentally friendly solution, but it works. These chemicals can create some potential risks in the final product and alleviating those risks through genetic engineering will benefit farmers and consumers alike.
Enzymatic browning is catalyzed by polyphenol oxidases (PPOs), which convert phenolic substrates to quinones. To reduce enzymatic browning, PPO activity must be lowered. In one experiment, the research team studied the editing of StPPO2 gene in the tetraploid cultivar Desiree. Specifically, the CRISPR/Cas9 system was applied in potatoes to induce mutations in the StPPO2 gene, responsible for the PPO activity and undesired enzyme content in potatoes. Selected lines with mutations within the four alleles of the target gene were grown and assayed for enzymatic browning and PPO activity levels in tubers. It was hypothesized that specific editing of this target gene would result in a lower PPO activity in the tuber, thus an overall reduction of the enzymatic browning.2
CRISPR/Cas9 And Its Off-Target Activity
Even though CRISPR/Cas9 can create targeted edits without introducing exogenous transgenes, there have been consistent issues with off-target activity, i.e. the introduction of unintended mutations. These problems have been reported using the CRISPR/Cas9 system in plants. Because of this clear issue, assaying all possible off-target mutations induced by the selected sgRNAs would only be possible through something like whole genome sequencing of the edited lines, a formidable goal with CRISPR/Cas9 – but not with Cas-CLOVER! Because Cas-CLOVER uses a dimeric system with a pair of guideRNAs, in contrast to CRISPR/Cas9’s single gRNA, the possibility of off-target mutations, and the need to backcross in order to remove those unintended mutations, all but disappears.
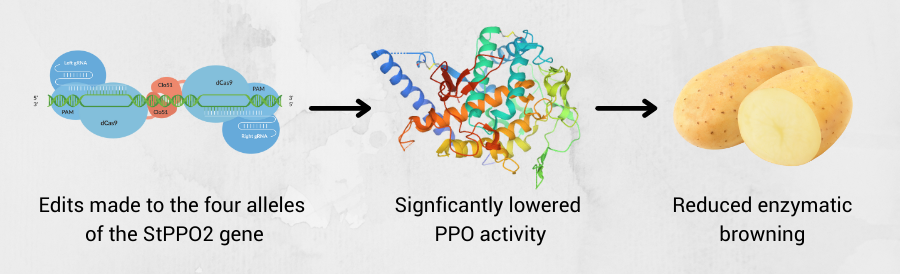
Figure 2: Using Cas-CLOVER to knockout the PPO gene can fix the fruit browning issue without off-target effects
Learn More About Cas-CLOVER And Demeetra
Cas-CLOVER’s distinctive dimeric gene editing technology proficiently minimizes off-target mutations, ensuring precise genetic modifications. If you’d like to learn more about how Demeetra can help accelerate your gene editing research with our toolkit, or to discuss licensing, get in contact with us by scheduling a call with us.
References
- Ackerman, J. (2015). Food: How Altered? National Geographic. Retrieved: https://www.nationalgeographic.com/environment/article/food-how-altered
- González MN, Massa GA, Andersson M, Turesson H, Olsson N, Fält A-S, Storani L, Décima Oneto CA, Hofvander P and Feingold SE (2020). Reduced Enzymatic Browning in Potato Tubers by Specific Editing of a Polyphenol Oxidase Gene via Ribonucleoprotein Complexes Delivery of the CRISPR/Cas9 System. Front. Plant Sci. 10:1649. doi: 10.3389/fpls.2019.01649
