The Complicated History of Wheat
Wheat has had a complicated history of dispersion, adaptation, and selection since it was domesticated in the Fertile Crescent between 8,000 and 10,000 years ago. As humans migrated from the African forests into savannah regions around 6 million years ago, grass species with small, hard seeds quickly entered the human diet. However, human development and increased grain consumption have gone hand-in-hand seemingly since the beginning. Consumption of cereal grains increased further with the advent of agriculture during the Neolithic era, and it has done so up until the present.
The fine, stretchy doughs and tasty white breads made with flour milled from wheat (Triticum aestivum L.) gained fame in Roman times. The expanded cultivation of wheat probably started in the Mediterranean region. Following two major human migration routes from the Fertile Crescent toward Europe and Asia via the Silk Road, bread wheat then differentiated in multiple new settings.

Even though the genetic structure of landraces are parallel to ancient human migration routes, data suggest a reshuffling through time related to breeding programs and the appearance of new alleles loaded with structural variations. Data suggests that recent wheat selection and spread has led to a modern germplasm that is highly unbalanced compared to the ancestral one found in landraces. Additionally, the genomes of wheat cultivars contain a sizable proportion of foreign DNA as a result of the introduction of wild relatives for crop enhancement.
Currently, 220 million acres of bread wheat are grown each year, yielding 700–750 million tons of grain that is used in a vast array of culinary goods. However, eating wheat is also linked to the emergence of a number of illnesses, such as allergies, auto-immune reactions, and non-celiac wheat sensitivity. Targeted gene editing, especially CRISPR/Cas9 and related gene editing technologies, are tools with considerable potential for plant development and breeding to address these illnesses and otherwise improve the crop for use as a food.
Gene Editing Advancements in Wheat Varieties
Whole-grain wheat-based goods that are safe for celiac disease are becoming more popular as eating whole-grain foods lowers the risk of many chronic diseases. However, conventional breeding alone cannot create wheat that is both celiac-safe and retains baking quality, due to the numerous gluten genes and overall complexity of the wheat genome. Within the context of food production, the use of gene editing for the development of wheat that is safe for celiacs should also be taken into consideration for further research (1).
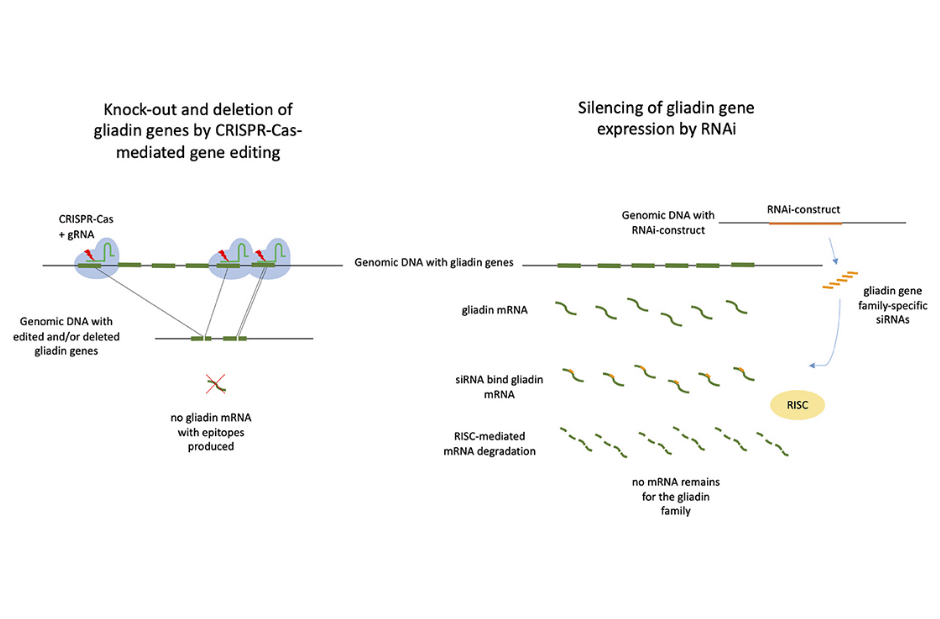
Many genome editing technologies are emerging as a potential way to fulfill a huge need in better crop breeding, and by developing a targeted strategy for generating novel genetic variation, we can begin to see better results. CRISPR/Cas-based genome editing has the ability to produce genetic variation in a targeted region of interest and it has the potential to alleviate some of the genetic bottlenecks imposed on modern crops. A variety of crop plants have already benefited from the use of CRISPR/Cas9, a gene editing technology that can substantially speed up breeding efforts with precise modifications in traits of interest (2).
As further research is drawn to the fields of genome editing and wheat biology, and as the number of gene editing platforms grows, more methods for precisely regulating gene expression in wheat will be made available. For example, scientists have created a strain of wheat that has been genetically altered to be resistant to fungal diseases without affecting the grain’s growth. Similar methods might also be effective for other commercial crops, such strawberries and cucumbers.
Various diseases originating from fungi, bacteria, and viruses all have the potential to significantly lower wheat yields and quality. To create wheat that is resistant to disease, disease-susceptibility genes are knocked out using CRISPR. In plants, loss of the function of mildew-resistance locus (MLO) confers broad-spectrum resistance to powdery mildew in plants. This makes MLO an ideal target for CRISPR (and other gene editing platforms) to enhance resistance to powdery mildew (3).
In barley, mutations in MLO cause stunted plants that typically yield up to 5% less grain than ordinary plants, which is an unacceptable deficit. In one wheat study, the research team produced the identical protective mutation in wheat’s six copies of the MLO gene using gene-editing techniques like CRISPR, and subsequently the gene-edited plants were successfully resistant to powdery mildew infection.
Deeper analysis of these altered plant’s genomes later revealed that substantial stretches of DNA on one chromosome, as well as a portion of the MLO gene were accidentally removed during editing. This led to a nearby gene called TMT3 becoming more active, which is somehow responsible for maintaining normal plant growth. Another point to mention is that although CRISPR/Cas9 can actually cleave sites, it also cleaves sites with a few mismatches to the target site “off-target mutations”, leading to downstream issues (4).
Sometimes, through many generations of back-crossing or crossing with wild-type plants, these potential off-target mutations can become eradicated, but not always due to limitations in chromosomal crossover during meiosis or “linkage drag”. By using alternative gene editing platforms that are more precise and have less off-targets than Cas9 these issues can be alleviated.
Challenges with CRISPR/Cas9-mediated Genome Editing in Wheat
Despite the fact that CRISPR/Cas9 applications have the potential to speed up agricultural improvement, there are a number of obstacles that prevent this ground-breaking technology from being fully utilized, especially in crops with large polyploid genomes.
One of the most important problems preventing the application of CRISPR/Cas9 technology in wheat is off-target gRNA binding and cleavage because of the existence of vast gene families and up to six homoeoalleles per gene. The effectiveness and specificity of the Cas9 nuclease are determined by the gRNA, making it a crucial part of the CRISPR/Cas9 system. High on-target activity and little off-target potential are characteristics of an efficient gRNA. In order to achieve maximum effectiveness and the highest targeting specificity for the specified genomic site, researchers may have to consider other strategies, like dual gRNA and dimeric nuclease systems, to improve targeting specificity.
Overall, results suggest that CRISPR genome editing system can easily be established on wheat protoplast and it has a huge potentiality for targeted manipulation of wheat genome for crop improvement purposes. However, this particular genome editing has also been a long-term challenge for continued research, particularly for plants possessing a complex genome. Even though the efficiency of CRISPR/Cas9 system has been shown with several studies from diploid plants, its application remains a challenge for plants with polyploid and complex genomes (5).
Another current barrier to the successful use of CRISPR technology to wheat improvement is the absence of reliable bioinformatics tools to allow the design of highly specific functional guide RNAs (gRNAs) and prediction of off-target sites in wheat. Additionally, a high dependence on in vitro culture and regeneration skills are reasons that genome editing technologies, such as CRISPR/Cas9, in wheat varieties are currently constrained. The callus culture and regeneration procedures required are typically time-consuming, expensive, and labor-intensive (6).
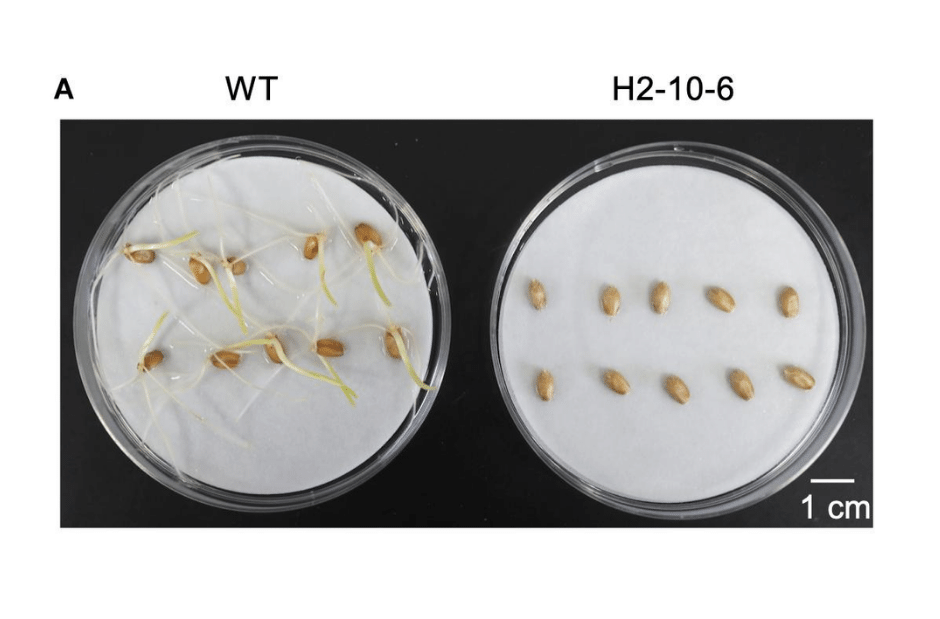
Figure 2: Effect of mutation of Haruyokoi TaQsd1 in seed dormancy. Only by knocking out all three homeologous TaQsd1 genes did researchers see this delay germination.
Furthermore, the use of genome editing in commercial varieties of important crops, like wheat, are severely constrained by the fact that these traditional culture-based transformation approaches are only relevant to genotypes that are amenable to cell culture and regeneration.
Future Prospects for Wheat Improvement Through Genome Editing
There are many existing challenges in wheat development and production, and the future heavily relies on precision gene modifications using advanced genome-editing technologies. Wheat will considerably benefit from genetic enhancement through the combination of genome editing and modern breeding techniques for sustainable global production. To enable wheat improvement through precise genome editing, additional research into various techniques to boost HDR efficiency are required.
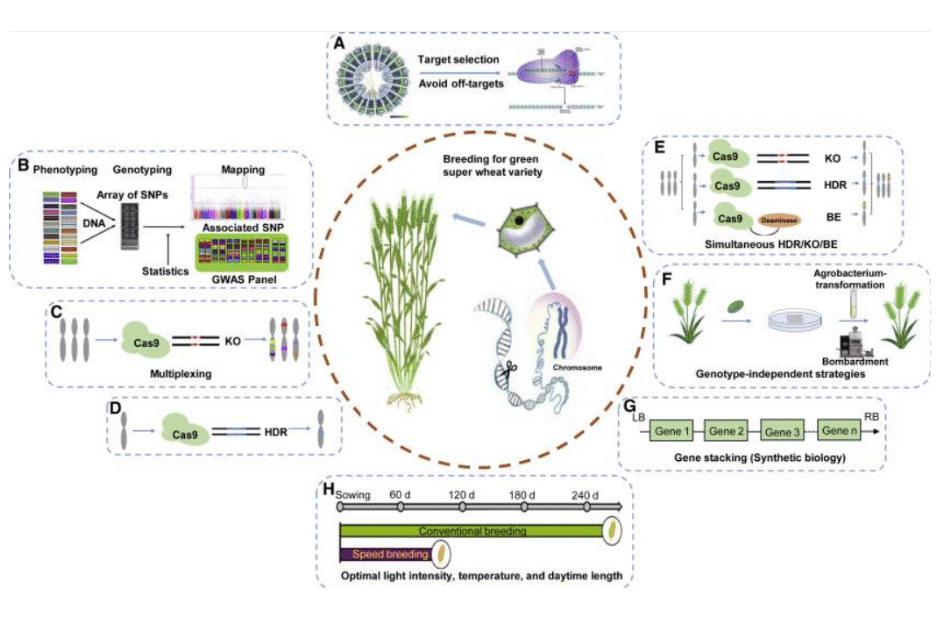
Figure 3: Breeding of a green super wheat variety through CRISPR/Cas-mediated gene editing and other breeding technologies.
The field of plant research has undergone quite a revolution, and CRISPR-mediated genome-editing tools now hold enormous promise for wheat biological research and genetic improvement. By developing various genome-editing toolkits, research teams will be able to target several desired alleles in commercial variety and improve wheat in a very efficient method,without compromising other prized traits without linkage drag from deleterious genes (7).
Cas-CLOVER is a Cleaner Gene Editing Alternative to CRISPR/Cas9
At Demeetra, we believe that the future of producing green super wheat varieties for sustainable agriculture will happen through advanced genome editing along with other cutting-edge breeding technologies. Despite significant progress in wheat genome editing, Cas-CLOVER technology may have more potential due to its dual gRNA structure and precise editing.
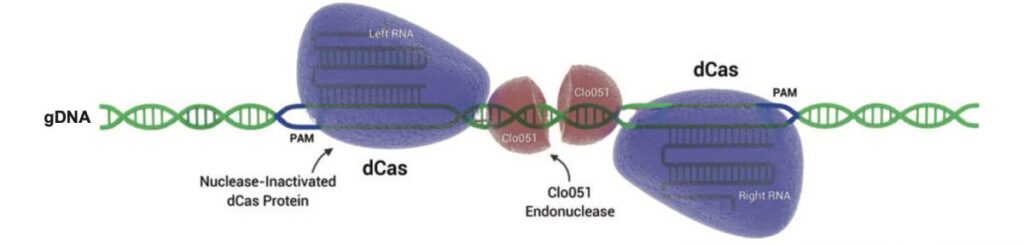
Fig 4: Cas-CLOVER is a gene editing system similar to CRISPR/Cas9 but uses unique advantages. It employs two gRNAs and dCas9 for DNA binding, with the Clo051 nuclease performing cleavage, ensuring greater precision and fewer off-target mutations. Cas-CLOVER also produces larger deletions for efficient gene knockouts and insertions, enhancing accuracy and establishing its own intellectual property separate from CRISPR/Cas9 patents.
Cas-CLOVER is a cleaner gene editing alternative to CRISPR/Cas9 for commercial applications. Similar indel frequencies can be achieved as compared with CRISPR/Cas9. Cas-CLOVER is a highly specific system demonstrating few detectable off-targets, which is very important for editing crops with highly repetitive and homologous genomes. We have validated Cas-CLOVER in plants by efficiently creating knockout mutations in the model organism and dicot tobacco, as well as monocot banana. Learn more about our more precise and easy-to-license Cas-CLOVER gene editing technology today.
References:
- Jouanin et al. (2020) CRISPR/Cas9 Gene Editing of Gluten in Wheat to Reduce Gluten Content and Exposure—Reviewing Methods to Screen for Coeliac Safety. Front. Nutr. 7:51. doi: 10.3389/fnut.2020.00051
- Lyzenga et al. (2021) Wheat improvement using genome editing technology. BioTechniques 2021 71:6, 577-579. https://doi.org/10.2144/btn-2021-0067
- Li et al. (2021) Recent advances in CRISPR/Cas9 and applications for wheat functional genomics and breeding. aBIOTECH.
- Stokstad E. Gene-edited wheat resists dreaded fungus without pesticides. Erik Stokstad, Science 2022. doi: 10.1126/science.ada1111
- Kim et al. (2018) CRISPR/Cas9 genome editing in wheat. Funct Integr Genomics. Jan;18(1):31-41. doi: 10.1007/s10142-017-0572-x.
- Liu et al. (2021) In planta Genome Editing in Commercial Wheat Varieties. Front. Plant Sci. 12:648841. doi: 10.3389/fpls.2021.648841
- Li et al (2021). Present and future prospects for wheat improvement through genome editing and advanced technologies. Plant Commun. 2021 Jun 5;2(4):100211. doi: 10.1016/j.xplc.2021.100211.
