The CRISPR-Cas9 system has revolutionized targeted genome engineering across a multitude of organisms, providing unprecedented simplicity in genetic manipulation. However, this technology is not without its limitations. Notable among these are the propensity for off-target mutagenesis, stringent licensing restrictions, and economically burdensome license terms.
In response to the limitations inherent to the single-guide RNA (sgRNA) Cas9 editing paradigm, scientists at Demeetra embarked on a research initiative to explore the viability of its proprietary Cas-CLOVER technology as a novel alternative for crop scientists. This endeavor was aimed at empirically validating the functional activity and efficacy of Cas-CLOVER's nucleic acid cleavage in tobacco, positioning it as a potentially more targeted and economically feasible genetic editing tool for agricultural biotechnology applications.
How Cas-CLOVER Works And How It Differs From Single-Guided Cas Gene Editing
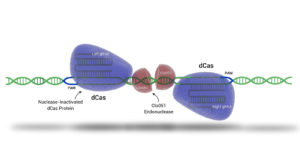
Cas-CLOVER is a patented dimeric gene editing system using a pair of guide RNAs (gRNAs), nuclease-inactivated proteins fused to our proprietary Clo051 endonuclease.
The fusion protein serves only as a linker between the guide pairs and Clo51; it is mutated and unable to cut DNA. Cleavage activity thus depends on the dimerization of the “obligate dimer” Clo51. The compositional and functional differences between Cas-CLOVER and CRISPR/Cas9 renders the system more precise than Cas9 and carves out independent intellectual property.
Demonstration of Cas-CLOVER Cutting Efficiency In Plants
Cas-CLOVER has demonstrated robust targeted mutagenesis in mammalian cells, including human T-cells, iPSCs and chinese hamster ovary (CHO) cells, but we wanted to open the door to Cas-CLOVER use in plants, trait discovery, and crop science applications.
Our strategy involved targeting the phytoene desaturase (PDS) gene in tobacco due to its visually pale and white phenotype. We focused the screening at the sequence level on two guide pair sites we called 1 and 4. Green, pale mixed, and white shoots were all screened to determine insertion and deletion “indel” efficiencies, a common measurement for genome editing. Then, we verified that only white and pale material was edited at an overall average of around 50%+ editing efficiency (white shoots/total shoots from plates).

Shoot phenotypes. Note the contrast between wild-type (green) from successfully manipulated PDS gene phenotype (white)
The full white phenotype shown here indicates all 4 alleles were knocked out with Cas-CLOVER. After fully optimizing our protocols we enhanced both the number of white/pale shoots per plate and genomic cutting efficiencies up to 90%+ as shown below.
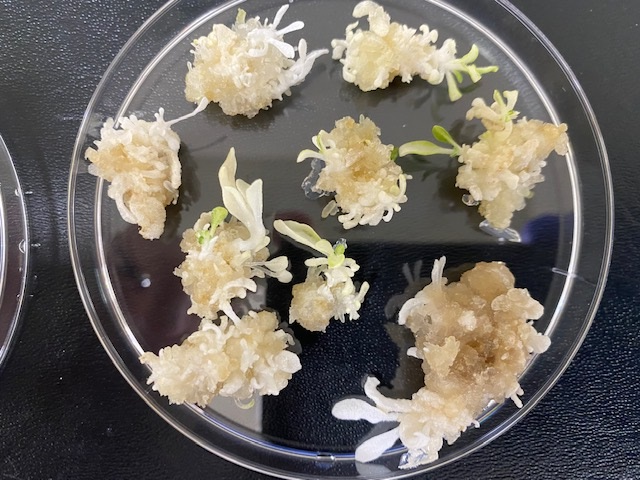
Phenotype of successfully edited tobacco shoots
Genetic modification using Cas-CLOVER was first confirmed by Next-Gen sequencing. Additionally, we analyzed individual transgenic plants using a combination of PCR, Sanger sequencing and applications like Synthego ICE shown below as indel percentages.
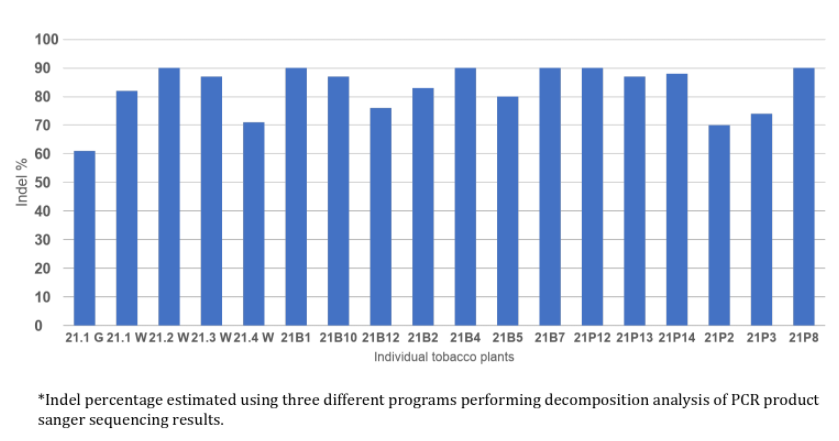
Estimated indel frequencies of individual plant subjects
Improved Downstream Processes For Cas-CLOVER Editing Through The plant T1 Generation
Eight (8) T0 edited plants were grown to the flowering stage, the plants were self-pollinated, and seeds were harvested. Following seed germination, seedlings were transferred to magenta boxes and grown large enough to check for edit stability in this T1 generation. Unpurified PCR products were screened using an optimum resolution on an automated DNA analysis machine, QIAxcel (<500bp size).
- Cas-CLOVER produces larger indels than other technologies
- The large indels are rapidly and easily detected by peak size & translated into a gel image

QIAxcel DNA analysis machine
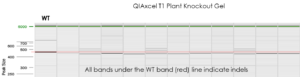
QIAxcel results demonstrating very high stability in the edited genome.
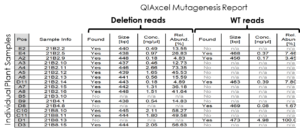
QIAxcel results, showing deletion versus wild type reads
T1 indels were then double checked using Synthego ICE. 14 of 16 samples sequenced and analyzed demonstrated 100% indel percentage. Two samples showed indel and WT sequences by both QIAxcel and ICE. Two other samples with WT peaks by QIAxcel revealed edited but smaller deletions by ICE not picked up in the QIAxcel screen.
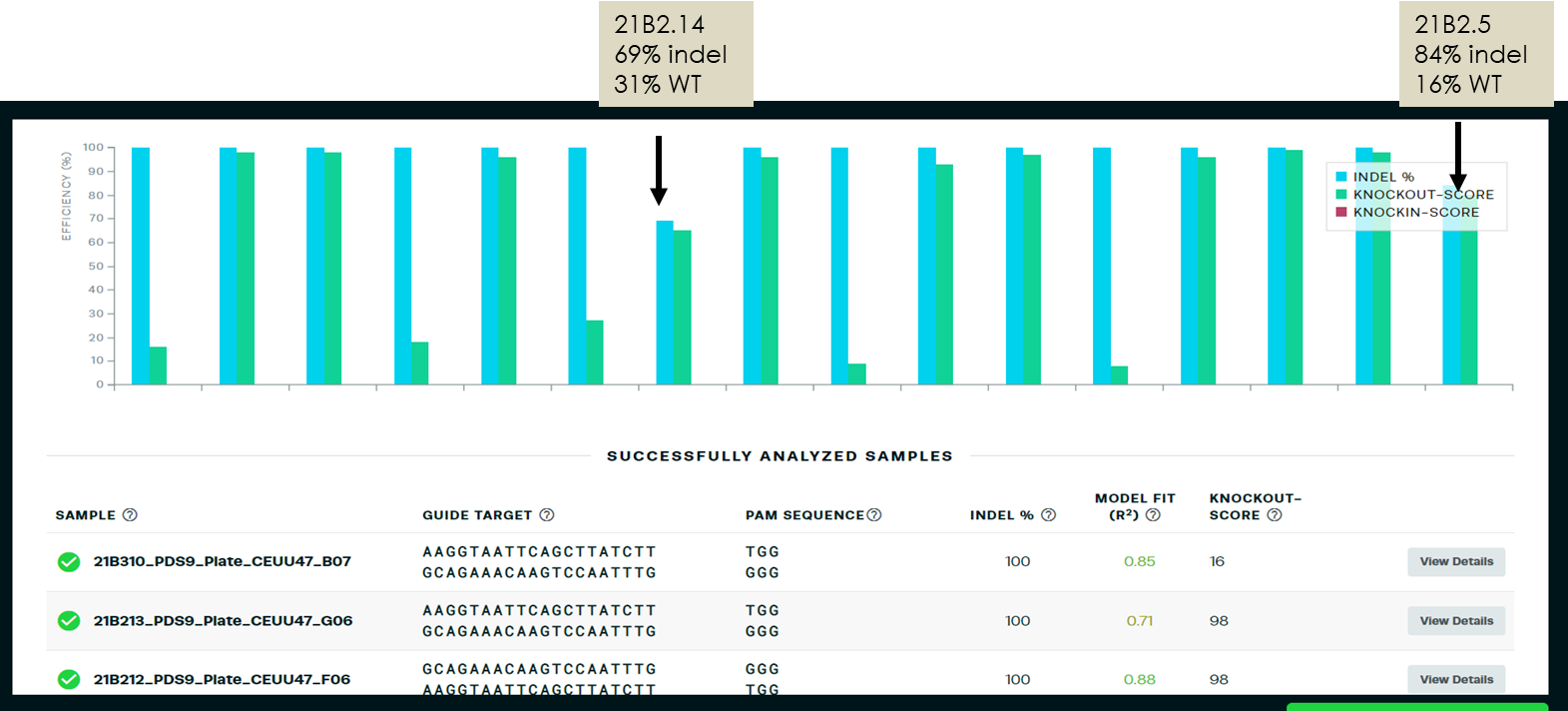
Synthego ICE results reflecting 100% indel for 14 of the 16 specimens and partial indel for the other 2
Demeetra’s Approach To Cas-CLOVER Licensing
This pioneering work conducted by Demeetra in refining the Cas-CLOVER technology presents a significant leap forward in the field of agricultural biotechnology, particularly in the realm of crop improvement. By addressing the limitations of the traditional CRISPR-Cas9 system, such as off-target effects and licensing constraints, Cas-CLOVER emerges as a highly precise, cost-effective, and efficient alternative for targeted genome engineering. Its unique dimeric gene editing mechanism, leveraging dual guide RNAs and the proprietary Clo051 endonuclease, has demonstrated substantial promise in plant model organism modifications, showcasing both high mutagenesis efficiency and edit stability through generations. This proof of concept establishes the versatile applicability of Cas-CLOVER across various plant species, offering new horizons for trait discovery, genetic improvement, and ultimately, the enhancement of crop resilience and productivity.
References
- Norman, D., Rector, K., Tateno, M., & Crawford, J. (2020). Targeted editing of Tobacco with Cas-CLOVER™: the clean alternative to CRISPR/Cas9 for plant genome editing. Plant Biology 2020. Demeetra AgBio, Lexington, KY