Parthenocarpy (the production of seedless fruit without prior fertilization) occurs in nature whether through a mutation that leads to negative consequences or as a benefit to shield against predators. It can also be exploited by scientists to achieve produce with longer shelf- lives or for practical purposes, such as large-scale sauce production, or more curb appeal to customers.
Think about the seedless watermelons you buy at your local grocery store. Seedless fruits are smaller and in the case with watermelons, this makes it easier to carry and perhaps not as wasteful if eating for one or two vs. the 4 to 8 people it takes to eat a traditional larger watermelon.
And, of course people appreciate the seedless option in regards to clean-up.
Gene editing technology allows for efficiently altering plant genomes in order to obtain the characteristics we desire which could be more practical and functional or weigh on the side of aesthetics and convenience.
Target Developmental Genes?
Traditionally, horticulturists breed plants by increasing the likelihood of producing certain genes as is done in the natural selection process. A gene of interest is noticed for a specific characteristic and then amplified in the population by breeding those plants with the genetic characteristic of interest.
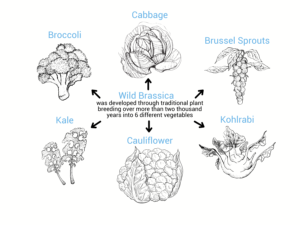
Figure 1: Traditional genetic engineering has done amazing things, but is inefficient and time-consuming
Nowadays, we can genetically alter in a more direct and precise manner. In doing so, it’s important to understand the signaling pathway and genetics involved to ensure the target gets altered without changing other functions. In the case of manipulation of fruit development, we look to the developmental genes and pathways involved.
The auxin response pathway regulates transcription in developmental genes; therefore, it’s the perfect pathway to develop direct gene editing tools to alter a plant’s seed production.
Altering Tomato Protein With Gene Editing Technology
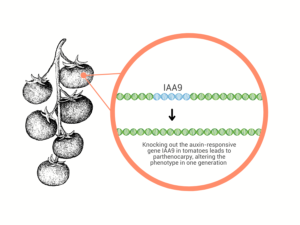
Figure 2: Gene editing technology can introduce desired traits in one generation, unlike traditional breeding
A recent study focused on using gene editing technology to alter the auxin-responsive protein, IAA9, in the tomato (Solanum lycopersicum) plant (SlIAA9). 1
Aux/IAA9 and auxin response factor 8 (ARF8) are involved in repressing fruit initiation without fertilization (parthenocarpy) in tomatoes. The researchers found that the SlIAA9 knock outs exhibited simple leaves compared to the compound leaves of the wild-types; shown in figure 1.
Additionally, fruit development was triggered before fertilization in those plants without SlIAA9, giving rise to parthenocarpy; also shown in figure 1.
Furthermore, the mutations were heritable in subsequent generations meaning it’s a sustainable genetic editing technique. 1
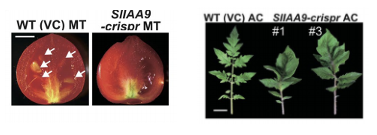
Figure 3: Tomato fruit with seeds (WT) and without (genetically modified fruit). Compound leaves in wild-type vs. simple leaves in the genetically modified leaves.
Novel Tools For Gene Editing In Plants
Previous research in fruit ripening enzymes used gene technology such as RNAi which reduces gene expression at the mRNA level while newer technologies such as CRISPR and Cas-CLOVER completely and permanently silences the gene at the DNA level. Novel gene editing proteins such as, Cas9, transcription activator-like effector nucleases (TALEN), and zinc-finger nucleases (ZFN) target specific genes in a deliberate manner. This is compared to previous tools that randomly insert, delete, or cause a frameshift in the sequence being edited; dramatically altering gene function.
Besides the temporary reduction vs. a permanent solution, RNAi silencing method suffers limitations due to high off-target effects. Off-targets in plants could lead to unwanted changes that reduce quality possibly leading to decreased yield, income, and /or health and regulatory concerns. The Cas-CLOVER gene editing system shown in Figure 2 utilizes a catalytically inactive Cas9 protein fused to the Clo51 nucleus domain which works as monomers recruited by a pair of guide RNAs (gRNA) to introduce targeted mutations. And when both subunits are properly recruited to the target-site, it leads to dimerization and activation of the clo51 nucleus domain, leading to targeted gene disruptions.
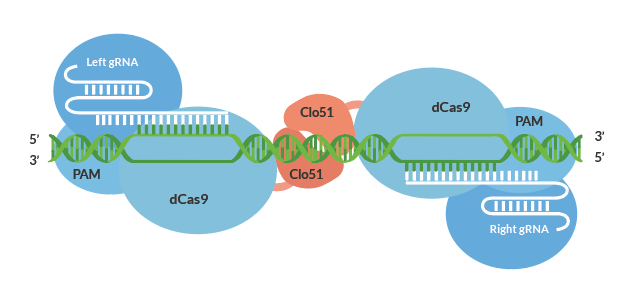
Figure 4: Cas-CLOVER gene editing system
Demeetra’s Cas-CLOVER Gene Editing Systems
Advanced gene editing technologies enable plant biotechnologists with the ability to make targeted knockouts and insertions. This scarless gene editing means no heterologous genes are left in the plant genome, producing a non-GMO crop and simplifying the process. We recognize the impact that gene editing tools have on the future of science, agriculture, and ultimately food preservation. In particular, we offer licenses for our novel gene editing Cas-CLOVER system.
Shown in a previous publication, we were able to validated the activity of Cas-CLOVER in plants by targeted inactivation of the RNA-dependent RNA polymerase 6 (RDR6) gene in tobacco achieving an efficiency of at least 18% in the first target which is significant for tobacco.
Enhancing tomato production is just one example that can benefit from gene editing technology. Our rapidly growing population with threats of climate change and global pandemics continue to be factors that pressure our food and medicine security, increasing the need for more productive agriculture traits and biomanufacturing systems is essential.
If you’d like to learn more about Cas-CLOVER, please contact us to schedule a meeting online.
References
Ueta, R., Abe, C., Watanabe, T., Sugano, S. S., Ishihara, R., Ezura, H., … Osakabe, K. (2017). Rapid breeding of parthenocarpic tomato plants using CRISPR/Cas9. Scientific Reports, 7(1), 1–8. https://doi.org/10.1038/s41598-017-00501-4
