In our modern society where we visit the grocery once a week versus daily, keeping produce fresh is a challenge. It’s a challenge worth investing in due to not only the economic value, but the nutritional importance as well.
Modern gene editing technology is allowing for many advantages such as longer lasting storage of produce; the tomato is a perfect example. Being able to keep tomatoes fresh for longer periods of time helps consumers save time and money. It’s a commonly used item in dishes all over the world; from curries and pasta sauce to being added fresh to salads and salsas.
Why Pectin?
The natural degradation of several plants’ (including tomatoes) cell wall decreases firmness which is good for taste since we want fresh, ripe produce; however, it’s obviously not great for transporting, and long term storage. Homogalacturonan (HG), rhamnogalacturonan-I (RG-I), and rhamnogalacturonan-II (RG-II) have been identified as pectin polysaccharides that are released from the cell wall during the ripening process through the breakage of covalent linkage. 1
Due to the role pectin plays in the texture of the tomato, it’s no wonder it has become the focus in gene editing systems in order to achieve longer-lasting firmness. In a recent study, researchers leveraged advanced DNA editing technologies to generate loss of function mutants to examine functionality in specific cell wall structural enzymes. 1
The scientists observed the effects on fruit softening and pectin localization in the cell wall (pericarp) of ripe fruit after after generating mutations in genes encoding the tomato pectin–degrading enzymes pectate lyase (PL), polygalacturonase 2a (PG2a), and b-galactanase (TBG4). 1
Their findings demonstrate the value in using gene editing systems. Specifically, they found that the three enzymes are necessary for normal ripening-related changes in cell-to-cell adhesion and that the loss of function led to a lower level of ripening-related activities.1
Ripening-related activities are measured in various ways. The three pectin degrading enzymes were found to have different functions.
Mutations in PL resulted in firmer fruits, but PG2a and TBG4 mutations altered color and weight; decreased color and increased size. Pectin localization, distribution, and solubility in the fruit cell wall are additional ripening-related measurements gathered in the study. Higher levels of deesterified homogalacturonan were found in the cell wall of the PL, PG2a, and TBG4 mutated cell lines compared to the control indicating less pectin degrading in the knockout cell lines.
These results are exciting for those interested in finding ways to increase storage and transportation times for tomatoes and other produce.
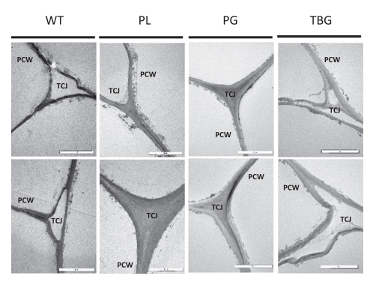
Sections cut from wild-type and the three mutated PL5, PG34, and TBG-8 lines were visualized under the transmission electron microscope and two representative micrographs shown for each line. The mutated fruits often appear larger than in other lines.
New Gene Editing Technologies Like CRISPR And Cas-CLOVER
Novel gene editing proteins such as, Cas9, transcription activator-like effector nucleases (TALEN), and zinc-finger nucleases (ZFN) target specific genes in a deliberate manner. This is compared to previous tools that randomly insert, delete, or cause a frameshift in the sequence being edited; dramatically altering gene function.
Previous research in fruit ripening enzymes used gene technology such as RNAi which reduces gene expression at the mRNA level while newer technologies such as CRISPR and Cas-CLOVER completely and permanently silences the gene at the DNA level and do not leave exogenous transgenes in the plant. Besides the temporary reduction versus a permanent solution, RNAi silencing method suffers limitations due to high off-target effects and GMO-status.
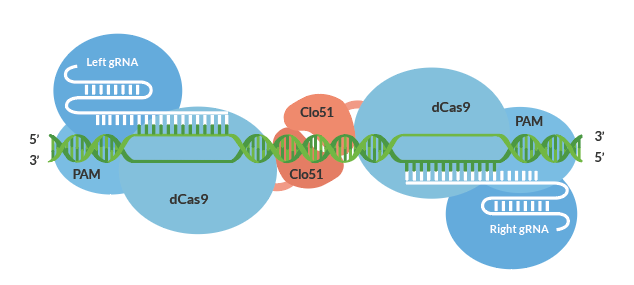
Cas-CLOVER was first introduced and validated to have little to no off-targets due to its dimeric nature in cell lines such as human T-cells². It utilizes a catalytically inactive Cas9 protein fused to the Clo51 nucleus domain which works as monomers recruited by a pair of guide RNAs (gRNA) to introduce targeted mutations. And when both subunits are properly recruited to the target-site, it leads to dimerization and activation of the Clo51 nucleus domain, leading to targeted gene disruptions.
Demeetra’s Advanced Gene Editing System Validated For The Plant Genome
Advanced gene editing technologies enable plant biotechnologists with the ability to make targeted knockouts and insertions. This scarless gene editing means no heterologous genes are left in the plant genome, producing a non-GMO crop and simplifying the process.
Shown in a previous publication, we validated the activity of Cas-CLOVER in plants by targeted inactivation of the RNA-dependent RNA polymerase 6 (RDR6) gene in tobacco achieving an efficiency of at least 18% in the first target which is significant for tobacco³.
Enhancing tomato production is just one example that can benefit from gene editing technology.
Our rapidly growing population with threats of climate change and global pandemics continue to be factors that pressure our food and medicine security, increasing the need for more productive agriculture traits and biomanufacturing systems is essential.
We recognize the impact that gene editing tools have on the future of science, agriculture, and ultimately food preservation. In particular, we offer licenses for their novel gene editing Cas-CLOVER system for commercial and academic research.
If you would like to learn more about the Cas-CLOVER system for gene editing in plant cells, please contact us here.
References
- Wang, D., Samsulrizal, N. H., Yan, C., Allcock, N. S., Craigon, J., Blanco-Ulate, B., … Seymour, G. B. (2019). Characterization of CRISPR mutants targeting genes modulating pectin degradation in ripening tomato 1[OPEN]. Plant Physiology, 179(2), 544–557. https://doi.org/10.1104/pp.18.01187
- Li, X., Wang, X., Tong, M., Tan, Y., Down, D., Shedlock, J., Ostertag, E.M., Poseida Therapeutics, Cas-CLOVER™: A High-Fidelity Genome Editing System for Safe and Efficient Modification of Cells for Immunotherapy https://demeetra.com/wp-content/uploads/2020/04/CRISPR-Congress-Poster-Cas-CLOVER-1.pdf
- Norman, D., Rector, K., Tateno, M., Crawford, J. “Targeted Editing of Tobacco with Cas-CLOVER the clean alternative to CRISPR/Cas9 for plant genome, Demeetra AgBio https://plantbiology2020.ipostersessions.com/Default.aspx?s=F7-2E-1F-D7-8F-B7-07-3F-C4-F2-8A-1C-5E-8B-08-F2